-Search query
-Search result
Showing 1 - 50 of 947 items for (author: thomas & dr)
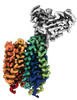
EMDB-43683: 
Cryo-EM structure of FLVCR2 in the inward-facing state with choline bound
Method: single particle / : Cater RJ, Mancia F
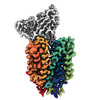
EMDB-43684: 
Cryo-EM structure of FLVCR2 in the outward-facing state with choline bound
Method: single particle / : Cater RJ, Mancia F

PDB-8vzn: 
Cryo-EM structure of FLVCR2 in the inward-facing state with choline bound
Method: single particle / : Cater RJ, Mancia F

PDB-8vzo: 
Cryo-EM structure of FLVCR2 in the outward-facing state with choline bound
Method: single particle / : Cater RJ, Mancia F

EMDB-19735: 
Full-length human cystathionine beta-synthase with C-terminal 6xHis-tag, basal state, helical reconstruction
Method: helical / : McCorvie TJ, Yue WW

EMDB-19736: 
Full-length human cystathionine beta-synthase with C-terminal 6xHis-tag, basal state, single particle reconstruction
Method: single particle / : McCorvie TJ, Yue WW

EMDB-19737: 
Full-length human cystathionine beta-synthase, basal state, helical reconstruction
Method: helical / : McCorvie TJ, Yue WW

EMDB-19738: 
Full-length human cystathionine beta-synthase, basal state, single particle reconstruction
Method: single particle / : McCorvie TJ, Yue WW

EMDB-19739: 
Full-length human cystathionine beta-synthase, basal state, partially degraded tetramer
Method: single particle / : McCorvie TJ, Yue WW
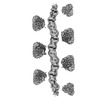
EMDB-19740: 
Full-length human cystathionine beta-synthase with C-terminal 6xHis-tag, SAM bound, activated state, helical reconstruction
Method: helical / : McCorvie TJ, Yue WW

EMDB-19741: 
Full-length human cystathionine beta-synthase with C-terminal 6xHis-tag, SAM bound, activated state, local helical reconstruction
Method: helical / : McCorvie TJ, Yue WW

EMDB-19742: 
Full-length human cystathionine beta-synthase with C-terminal 6xHis-tag, SAM bound, activated state, local single particle reconstruction
Method: single particle / : McCorvie TJ, Yue WW

PDB-8s5h: 
Full-length human cystathionine beta-synthase with C-terminal 6xHis-tag, basal state, helical reconstruction
Method: helical / : McCorvie TJ, Yue WW

PDB-8s5i: 
Full-length human cystathionine beta-synthase with C-terminal 6xHis-tag, basal state, single particle reconstruction
Method: single particle / : McCorvie TJ, Yue WW

PDB-8s5j: 
Full-length human cystathionine beta-synthase, basal state, helical reconstruction
Method: helical / : McCorvie TJ, Yue WW

PDB-8s5k: 
Full-length human cystathionine beta-synthase, basal state, single particle reconstruction
Method: single particle / : McCorvie TJ, Yue WW

PDB-8s5l: 
Full-length human cystathionine beta-synthase, basal state, partially degraded tetramer
Method: single particle / : McCorvie TJ, Yue WW

PDB-8s5m: 
Full-length human cystathionine beta-synthase with C-terminal 6xHis-tag, SAM bound, activated state, helical reconstruction
Method: helical / : McCorvie TJ, Yue WW

EMDB-15525: 
Cryo-EM structure of the RecA postsynaptic filament from S. pneumoniae
Method: helical / : Perry TN, Fronzes R, Polard P, Hertzog M

PDB-8amf: 
Cryo-EM structure of the RecA postsynaptic filament from S. pneumoniae
Method: helical / : Perry TN, Fronzes R, Polard P, Hertzog M

EMDB-41374: 
Antibody N3-1 bound to RBDs in the up and down conformations
Method: single particle / : Hsieh CL, McLellan JS
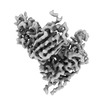
EMDB-41382: 
Antibody N3-1 bound to RBD in the up conformation
Method: single particle / : Hsieh CL, McLellan JS
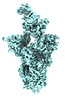
EMDB-41399: 
Antibody N3-1 bound to SARS-CoV-2 spike
Method: single particle / : Hsieh CL, McLellan JS

PDB-8tm1: 
Antibody N3-1 bound to RBDs in the up and down conformations
Method: single particle / : Hsieh CL, McLellan JS

PDB-8tma: 
Antibody N3-1 bound to RBD in the up conformation
Method: single particle / : Hsieh CL, McLellan JS
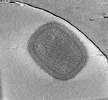
EMDB-18916: 
Cryotomogram of mature Vaccinia virus (WR) virion
Method: electron tomography / : Calcraft T, Hernandez-Gonzalez M, Nans A, Rosenthal PB, Way M

EMDB-18917: 
Subtomogram average of the Vaccinia virus (WR) portal complex in mature virions
Method: subtomogram averaging / : Calcraft T, Hernandez-Gonzalez M, Nans A, Rosenthal PB, Way M

EMDB-18918: 
Subtomogram average of the Vaccinia virus (WR) A4/A10 palisade trimer in mature virions
Method: subtomogram averaging / : Calcraft T, Hernandez-Gonzalez M, Nans A, Rosenthal PB, Way M

PDB-8r5i: 
In situ structure of the Vaccinia virus (WR) A4/A10 palisade trimer in mature virions by flexible fitting into a cryoET map
Method: subtomogram averaging / : Calcraft T, Hernandez-Gonzalez M, Nans A, Rosenthal PB, Way M

EMDB-40856: 
Single particle reconstruction of the human LINE-1 ORF2p without substrate (apo)
Method: single particle / : van Eeuwen T, Taylor MS, Rout MP
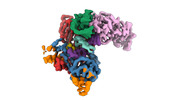
EMDB-40858: 
Structure of LINE-1 ORF2p with template:primer hybrid
Method: single particle / : van Eeuwen T, Taylor MS, Rout MP
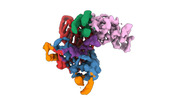
EMDB-40859: 
Structure of LINE-1 ORF2p with an oligo(A) template
Method: single particle / : van Eeuwen T, Taylor MS, Rout MP

PDB-8sxt: 
Structure of LINE-1 ORF2p with template:primer hybrid
Method: single particle / : van Eeuwen T, Taylor MS, Rout MP

PDB-8sxu: 
Structure of LINE-1 ORF2p with an oligo(A) template
Method: single particle / : van Eeuwen T, Taylor MS, Rout MP

EMDB-29400: 
Acinetobacter baylyi LptB2FG bound to lipopolysaccharide and a macrocyclic peptide
Method: single particle / : Pahil KS, Gilman MSA, Kruse AC, Kahne D

EMDB-29401: 
Acinetobacter baylyi LptB2FG bound to lipopolysaccharide.
Method: single particle / : Pahil KS, Gilman MSA, Kruse AC, Kahne D

EMDB-29402: 
Acinetobacter baylyi LptB2FG bound to lipopolysaccharide and RG6006
Method: single particle / : Pahil KS, Gilman MSA, Kruse AC, Kahne D

EMDB-29403: 
Acinetobacter baylyi LptB2FG bound to lipopolysaccharide and a macrocyclic peptide
Method: single particle / : Pahil KS, Gilman MSA, Kruse AC, Kahne D

EMDB-29404: 
Acinetobacter baylyi LptB2FGC
Method: single particle / : Pahil KS, Gilman MSA, Kruse AC, Kahne D

EMDB-42206: 
Acinetobacter baylyi LptB2FG bound to Acinetobacter baylyi lipopolysaccharide
Method: single particle / : Pahil KS, Gilman MSA, Baidin V, Kruse AC, Kahne D

EMDB-42207: 
Acinetobacter baylyi LptB2FG bound to Acinetobacter baylyi lipopolysaccharide and a macrocyclic peptide
Method: single particle / : Pahil KS, Gilman MSA, Baidin V, Kruse AC, Kahne D

PDB-8frl: 
Acinetobacter baylyi LptB2FG bound to lipopolysaccharide and a macrocyclic peptide
Method: single particle / : Pahil KS, Gilman MSA, Kruse AC, Kahne D

PDB-8frm: 
Acinetobacter baylyi LptB2FG bound to lipopolysaccharide.
Method: single particle / : Pahil KS, Gilman MSA, Kruse AC, Kahne D

PDB-8frn: 
Acinetobacter baylyi LptB2FG bound to lipopolysaccharide and Zosurabalpin
Method: single particle / : Pahil KS, Gilman MSA, Kruse AC, Kahne D

PDB-8fro: 
Acinetobacter baylyi LptB2FG bound to lipopolysaccharide and a macrocyclic peptide
Method: single particle / : Pahil KS, Gilman MSA, Kruse AC, Kahne D

PDB-8frp: 
Acinetobacter baylyi LptB2FGC
Method: single particle / : Pahil KS, Gilman MSA, Kruse AC, Kahne D

PDB-8ufg: 
Acinetobacter baylyi LptB2FG bound to Acinetobacter baylyi lipopolysaccharide
Method: single particle / : Pahil KS, Gilman MSA, Baidin V, Kruse AC, Kahne D

PDB-8ufh: 
Acinetobacter baylyi LptB2FG bound to Acinetobacter baylyi lipopolysaccharide and a macrocyclic peptide
Method: single particle / : Pahil KS, Gilman MSA, Baidin V, Kruse AC, Kahne D

EMDB-40969: 
Uncrosslinked nNOS-CaM oxygenase homodimer
Method: single particle / : Lee K, Pospiech TH, Southworth D

EMDB-40970: 
DSBU crosslinked nNOS-CaM oxygenase homodimer
Method: single particle / : Lee K, Pospiech TH, Southworth D
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
